How are we watching for the next COVID variant?
Sequencing efforts have improved, but experts say there's a long way to go.
This is a MedPage story.
The only certain thing about the future of SARS-CoV-2 variants is that nothing is certain -- but researchers in the U.S. are doing their best to keep an eye out for the next troublesome variant, even in the face of numerous challenges.
While the systems in place to identify and track new variants have come a long way, Dr. Stephen Morse, an epidemiologist at Columbia University's Mailman School of Public Health, told MedPage Today that "for the first time we have a really fairly detailed, not complete, but fairly detailed picture of how this is unfolding."
A key variant tracker is GISAID, which started as a way to track influenza but grew during the pandemic into a global database where almost anyone can upload COVID-19 sequencing data. It's been behind the discovery of new strains, and it's by far the largest repository of genetic data on the virus.
"People ... found some of these new sequences simply by looking at how people had uploaded [to GISAID], so that's kind of a reactive approach," said Morse, noting that more institutions are now actively sampling, and the Centers for Disease Control and Prevention "is now testing and sequencing, or collecting sequences."
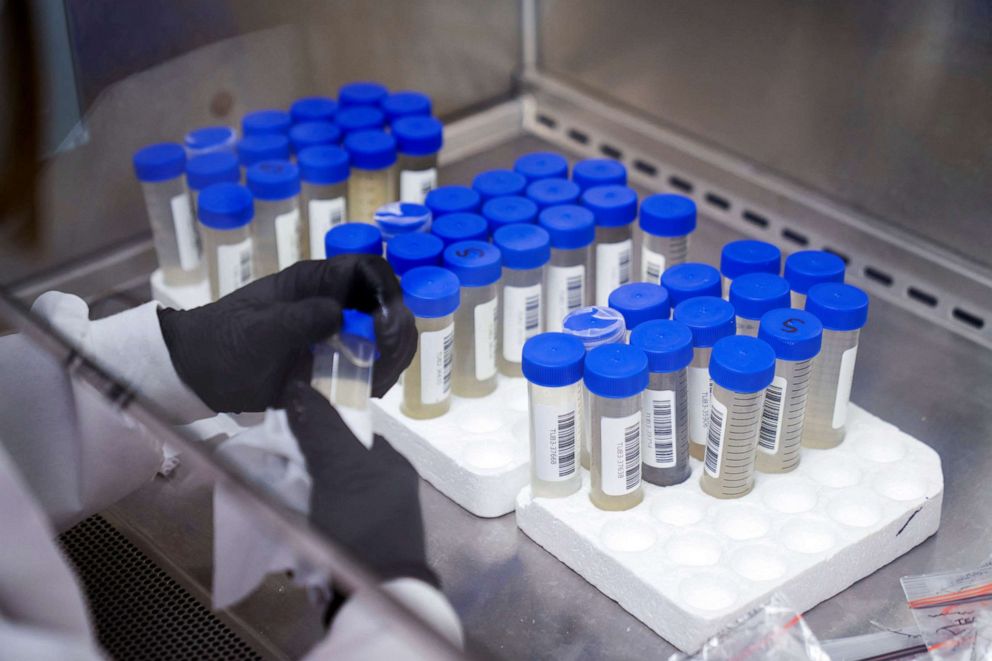
The CDC is collecting sequences through a program launched in November 2020 called the National SARS-CoV-2 Strain Surveillance program. Initially, the project only asked for 10 samples from each state every two weeks, but is now sequencing many more than its initial goal of 750 samples per week.
Unlike GISAID, NS3 is meant to be nationally representative. The CDC asks states to send samples that represent a variety of demographics, locations, and clinical characteristics to reflect the makeup of the country as a whole -- but some states send more or less than their proportional share of the samples.
"We go back to when it started, but we could have done a lot more early on," Morse said. But "When you're in the middle of a wildfire, you don't have really time to sort of develop or combine a system. You go and fight the wildfire with the systems you can get."
Just having whole genome data on variants of interest isn't enough. "The issue with the sequencing is, you need more than just the sequencing, you need to be able to link the sequence to the cases," said Dr. Sharon Balter, director of the Acute Communicable Disease Control and Prevention Division of the Los Angeles County Department of Public Health.
Public health departments rely on a web of players to connect the dots between sequenced samples and outcomes, which lead to policy decisions. Departments look at the GISAID data for their state or area, and watch other countries that have a more robust sampling approach. But some also do their own sequencing, or have help doing it.
In Los Angeles County, for example, Balter said universities and commercial labs do most of the sequencing, and then report their results to the county. If the person whose sample was sequenced ends up being hospitalized, public health officials must link their sequence to a personal identifier and get data from the hospital on their admission status, ICU status, intubation status, or fatality.
They also link this data with their vaccine registry to find out if the person was vaccinated or not. This way, the department can see if one variant is gaining traction in a particular area, or leading to more hospitalizations and can take action accordingly.
Wastewater monitoring
Environmental surveillance also has emerged as a way to get a broader picture of the pandemic's trajectory. Dr. Kartik Chandran, a professor of earth and environmental engineering at Columbia University, leads a project that conducts wastewater surveillance for the university and for Bergen County, New Jersey.
For any given sampling area, "if performed properly, that surveillance could give us a good imprint of that corresponding population -- every day, of the entire population," Chandran said. "And so that's the big advantage here."
His team collects samples from buildings and can sequence genetic material from COVID-19 present in infected people's gastrointestinal tract. He's watched as various variants have battled it out over the last two years. And unlike surveillance that relies on testing individuals, Chandran's team can get a snapshot of the overall COVID-19 concentration very quickly.
"Our turnaround times for a fairly significant number of samples is a few hours now," he said. "To my knowledge, even commercial labs can't even come close to this." But genomic sequencing for a breakdown of particular variants takes longer than getting the concentration data, and, Balter noted, it can't be linked back to particular cases or individuals.

The New York Department of Environmental Protection also tests wastewater from 14 sites at "sewersheds" or wastewater facilities to get a picture of city-wide trends in COVID-19 concentration. The DEP also shared data on a portion of the variants they were able to sequence (the most "clinically abundant" like alpha, delta, and omicron) with the city.
This was part of a pilot program that found wastewater variant data was consistent with city testing data. According to an email from New York State Department of Health spokesperson Erin Silk, the program "is being expanded to cover all counties and to include sequencing for the analysis of COVID-19 variants" and "will use high-speed sequencing methods to facilitate the rapid detection and identification of variants and their circulation throughout the State."
Limitations of surveillance
As the tools to monitor and track variants have evolved, their shortcomings have been revealed, experts said.
"These systems weren't necessarily existing, thriving, or necessarily set up for this kind of a job," Dr. Bryan Lewis of the University of Virginia's Biocomplexity Institute, said of sequencing before the pandemic. "It was more of a research project, so timeliness wasn't important."
The ability to track specific variants through sequencing -- and send samples and data to the CDC -- still varies from one level of government to the next, and depends largely on the resources a locality or state has at their disposal.
"In a perfect world, every jurisdiction would have the resources to do this," said Balter. "But we don't live in a perfect world, as you know."
Equipment required for whole genome sequencing is expensive, and public health departments may not have the capacity to build and run labs themselves -- or even outsource the job.
Then there's the lag time for sequencing itself. Balter said it takes 10 days from the time they have the isolate or sample to get the full sequence. Despite technological advancements, making sense of a virus with 30,000 base pairs is no easy feat.
What's more, the links between local, state, and federal systems need time to crystallize. Tying together data from sequencing labs, hospitals, and vaccine registries even on a local level -- what Balter called "linkages" -- isn't easy.
"It's a very time-consuming process, because those are all independent systems," she said. Health data from states must be de-personalized for the federal NS3 program, Balter said, adding another logistical hurdle.
A lack of a system robust enough to collect, sequence, and share data on variants when COVID-19 hit led to what Lewis described as a piecemeal approach, especially with GISAID.
"You get these sort of like ad hoc little side projects that people are doing ... to try and contribute to the global effort," he said. "So that's another part of this not-comprehensive, ad hoc approach."
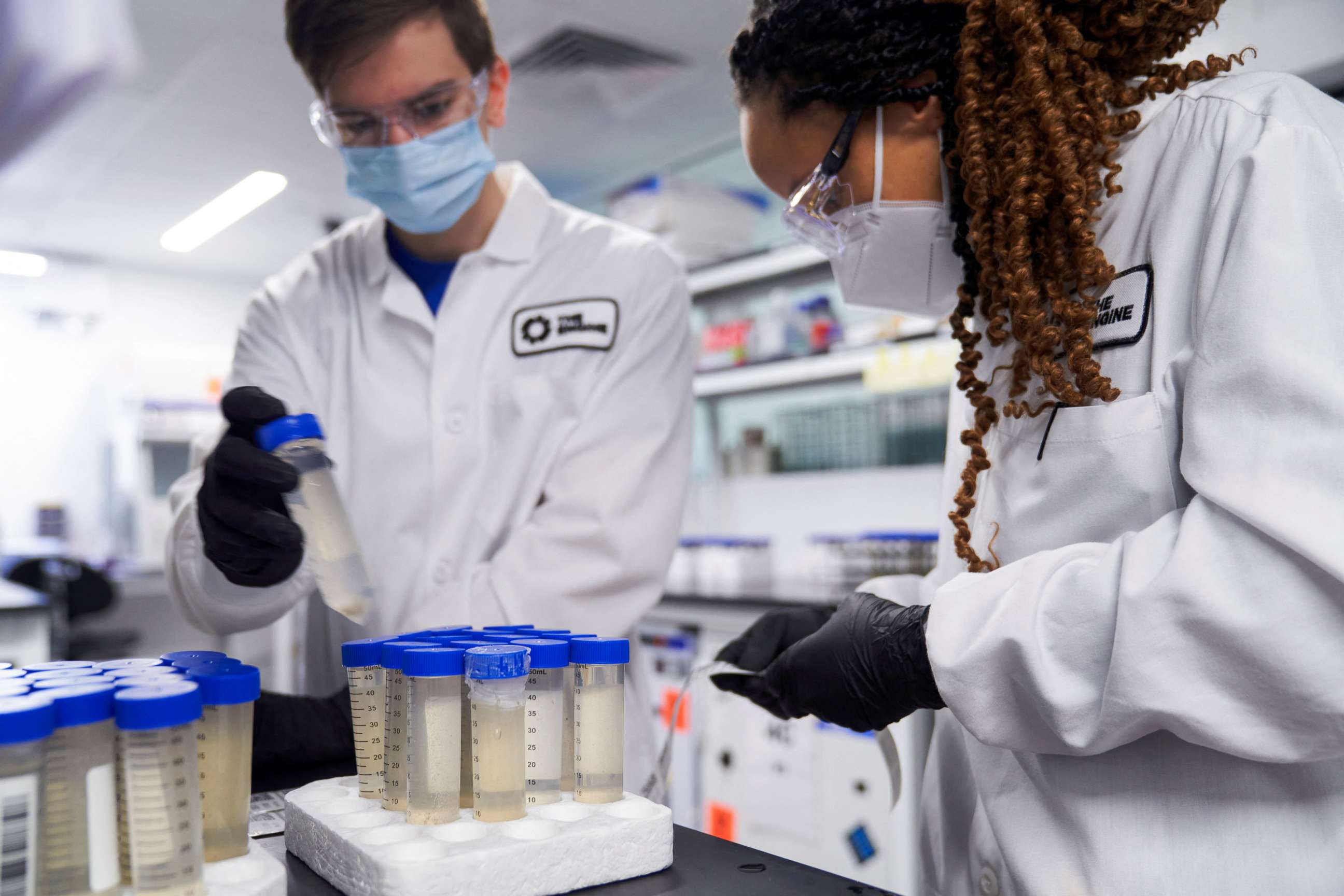

Representative sampling, which is what the CDC's sequencing surveillance system attempts to do, has also proven a difficult task. Collections of sequences sent or uploaded should ideally reflect the actual distribution of a population's characteristics.
"[It] would have been nice if we had a plan kind of like this started a year earlier than [COVID] came into fruition," Lewis said. "That would be very useful."
This lag is why the U.S. is so reliant on data from other countries to predict the path of variants here.
"In essence, Denmark and the U.K. can basically sequence all the cases that they have," Lewis said. "The United States isn't set up to do that. We don't spend the money and the resources."
Of the U.S. samples in GISAID, for example, some may have been submitted from a pool of adults over 65, or some specifically from patients with autoimmune disorders, which can end up over- or under-representing some populations.
In the meantime, experts say that they hope genomic surveillance systems can be used in the future, either to detect new threats or to make sense of what exactly happened with COVID-19. Knowing if a particular sequence, for example, predicts long COVID, Lewis said, isn't out of the question.
"This is a pretty unprecedented event," he said, "and it would be good for us to learn as much as possible about this."




